Structure and function of signaling complexes
The Biomolecular Interactions Research Group studies protein-protein and protein-drug interactions using biophysical/biochemical techniques. There is a strong interest in systems biology, and the analysis spans a wide spectrum: from atomic resolution structures to cellular behavior. The main focus is on complexes involved in cellular signaling. We use in vitro techniques such as three-dimensional structure determination, protein-protein interaction and enzymatic activity measurements as well as cell based assays. These mechanistic studies are often complemented by large-scale experimental approaches and systems-level computational simulations. The goal of the systems-level studies is the identification of new protein-protein complexes through which the cell’s signaling machinery could be specifically targeted to affect cell growth and death. Our mission is the discovery of new pharmaceutical strategies to combat cancer and chronic inflammation. We design new drugs by screening compound libraries and develop hit compounds through structure based rational design.
Protein kinases in cell signaling
Lots of diseases (e.g. cancer, chronic inflammation) emerge due to some mechanistic malfunctions in signaling networks. Our cells are subjected to diverse extracellular stimuli and they need to respond accordingly to complex patterns of inputs. The sensing of millions of these extracellular inputs, intracellular processing, and triggering the activation of the correct downstream effector proteins are all done by a heavily interconnected protein network. The effector proteins may be metabolic enzymes, transcription factors, or proteins involved in cell movements, which are regulated by protein phosphorylation. The latter is carried out by proteins kinases, and these enzymes also bind to each other and build kinase cascades. The intracellular signaling pathways controlling cell growth and death are “coming together” via complexes of protein kinases.
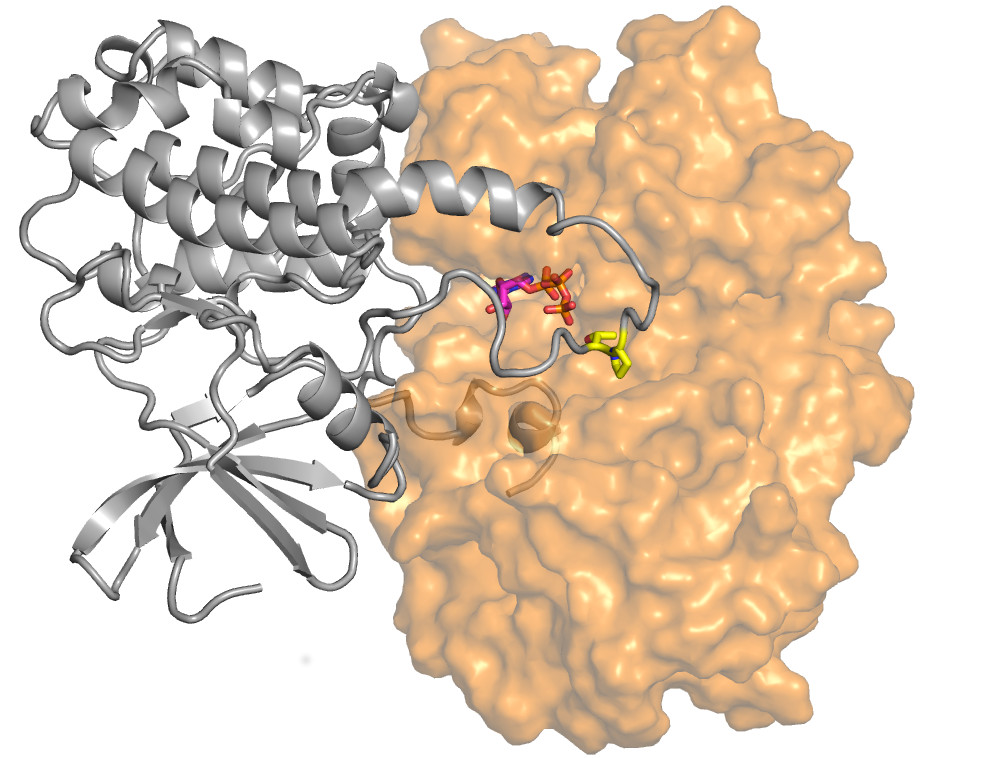
The portrait of a molecular switch involved in promoting cellular growth. The figure shows the crystal structure of the ERK2-RSK1 protein kinase complex (ERK2 in orange and RSK1 in gray). The yellow segment in the gray molecule works as a molecular switch, if it is modified by the orange molecule then the former gets turned on, and this ultimately will promote growth in cells.
Protein kinases are often targeted by proteins from pathogens and we can learn a great deal on how protein kinase-based cellular networks could be artificially manipulated by studying these proteins. The ORF45 protein from Kaposi’s sarcoma herpesvirus (KSHV) binds to the active ERK-RSK complex and promotes cell growth. After exploring the molecular logic of protein-protein binding within the ternary complex and by carrying out computational simulations based on a mechanistically correct model including other interaction partners, we can understand how the viral protein hijacks this important complex for the up-regulation of a cell growth promoting signaling pathway. ORF45 “attacks” pivotal protein-protein surfaces on ERK and RSK by short linear motifs that are also used to make physical links to the enzyme’s natural protein partners (e.g. upstream kinases, phosphataes, and substrate). It is interesting that the same surfaces (or interaction pockets) are also targeted by proteins from the plague bacterium or from the encephalomyelitis virus.
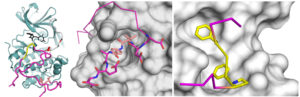
The tricks of a viral protein targeting the ERK-RSK signaling complex. The crystal structure of the ORF45 protein from Kaposi’s sarcoma (magenta) and RSK2 kinase (teal), and the molecular image of the way how the viral protein “attacks” two cell growth controlling enzymes (RSK2 and ERK2, shown in gray surface representation) by short peptide motifs (salmon and yellow, respectively).
Small molecules affecting protein-protein interactions
Current strategies aim to affect cell signaling by directly inhibiting the activity of protein kinases. The modulation of protein kinase function, however, could also be achieved by targeting specific kinase-based protein-protein complexes. Our goal is the synthesis of new drugs that indirectly affect the activity of selected protein kinases, for example by stabilizing inactive complexes or by interfering with an activating protein-protein interaction.
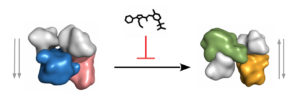
The 3D structure of a central protein kinase heterodimer involved in inflammation and its inhibition by a complex-specific inhibitor. The p38-MK2 complex can adopt two different quaternary structures: 1) inactive (left), which contains non-phosphorylated p38 (salmon) and it displays a parallel conformation regarding the relative orientation of the two kinase domains, 2) anti-parallel active (right), containing phosphorylated p38 (orange). The N-terminal lobes of the kinase domains are colored in gray, while the surface of the C-terminal lobes are colored differently. An anti-inflammatory drug blocks the natural “transition” between the two states of the p38-MK2 complex because it prefers binding to the inactive complex. This compound prevents the activation of the pro-inflammatory enzyme, MK2 (blue/green), by stabilizing the structure of the inactive p38-MK2 complex.
Publications:
Fontosabb publikációk (2012-2022):
Alexa A, Sok P, Gross F, Albert K, Kobori E, Póti ÁL, Gógl G, Bento I, Kuang E, Taylor SS, Zhu F, Ciliberto A, Reményi A.
Nat Commun. 2022 Jan 25;13(1):472. doi: 10.1038/s41467-022-28109-x
Co-regulation of the transcription controlling ATF2 phosphoswitch by JNK and p38.
Kirsch K, Zeke A, Tőke O, Sok P, Sethi A, Sebő A, Kumar GS, Egri P, Póti ÁL, Gooley P, Peti W, Bento I, Alexa A, Reményi A.
Nat Commun. 2020 Nov 13;11(1):5769. doi: 10.1038/s41467-020-19582-3
MAP Kinase-Mediated Activation of RSK1 and MK2 Substrate Kinases.
Sok P, Gógl G, Kumar GS, Alexa A, Singh N, Kirsch K, Sebő A, Drahos L, Gáspári Z, Peti W,Reményi A.
Structure. 2020 Oct 6;28(10):1101-1113.e5. doi: 10.1016/j.str.2020.06.007
Disordered Protein Kinase Regions in Regulation of Kinase Domain Cores.
Gógl G, Kornev AP, Reményi A*, Taylor SS*
Trends Biochem Sci. 2019 Apr;44(4):300-311. doi: 10.1016/j.tibs.2018.12.002.
Dynamic control of RSK complexes by phosphoswitch-based regulation.
Gógl G, Biri-Kovács B, Póti ÁL, Vadászi H, Szeder B, Bodor A, Schlosser G, Ács A, Turiák L, Buday L, Alexa A, Nyitray L, Reményi A.
FEBS J. 2018 Jan;285(1):46-71. doi: 10.1111/febs.14311
Zeke A, Bastys T, Alexa A, Garai Á, Mészáros B, Kirsch K, Dosztányi Z, Kalinina OV, Reményi A.
Mol Syst Biol. 2015 Nov 3;11(11):837. doi: 10.15252/msb.20156269
Gógl G, Schneider KD, Yeh BJ, Alam N, Nguyen Ba AN, Moses AM, Hetényi C, Reményi A*, Weiss EL*
PLoS Biol. 2015 May 12;13(5):e1002146. doi: 10.1371/journal.pbio.1002146
Structural assembly of the signaling competent ERK2-RSK1 heterodimeric protein kinase complex.
Alexa A, Gógl G, Glatz G, Garai Á, Zeke A, Varga J, Dudás E, Jeszenői N, Bodor A, Hetényi C, Reményi A.
Proc Natl Acad Sci U S A. 2015 Mar 3;112(9):2711-6. doi: 10.1073/pnas.1417571112.
Glatz G, Gógl G, Alexa A, Reményi A.
J Biol Chem. 2013 Mar 22;288(12):8596-8609. doi: 10.1074/jbc.M113.452235
Specificity of linear motifs that bind to a common mitogen-activated protein kinase docking groove.
Garai Á, Zeke A, Gógl G, Törő I, Fördős F, Blankenburg H, Bárkai T, Varga J, Alexa A, Emig D, Albrecht M, Reményi A.
Sci Signal. 2012 Oct 9;5(245):ra74. doi: 10.1126/scisignal.2003004
International collaborations:
Susan S. Taylor – University of California – San Diego, USA
Andrea Ciliberto – IFOM, Milánó, Olaszország
Eric L. Weiss – Northwestern University, USA
Marie Bogoyevitch – University of Melbourne, Ausztrália
Krishna Rajalingam – Johannes Gutenberg-Universität Mainz, Németország
EU-OPENSCREEN: http://www.eu-openscreen.eu/
Leader
Attila Reményi
Alumni
MSc
Ágnes Szonja Garai (ELTE biology)
Gergő Gógl (ELTE chemistry)
Ferenc Förős (ELTE biology)
Martina Rádli (ELTE biology)
Gábor Glatz (ELTE biology)
Boglárka Zámbó (ELTE biology)
Sarolt Magyary (ELTE chemistry)
Evelin Németh (ELTE chemistry)
Tamás Takács (ELTE biology)
Orsolya Ember (ELTE chemistry)
Bettina Balázs (SE medicine)
PhD
Gábor Glatz
Ágnes Szonja Garai
Gergő Gógl
András Zeke
Klára Kirsch
Sing Neha
Post-doc
Imre Törő
Domonkos Bartis
Krisztina Paál
András Zeke
Eszter Szarka Kállainé
Péter Egri